The characterization of a new set of EST-derived simple sequence repeat (SSR) markers as a resource for the genetic analysis of Phaseolus vulgaris – topic of research paper in Biological sciences. Download
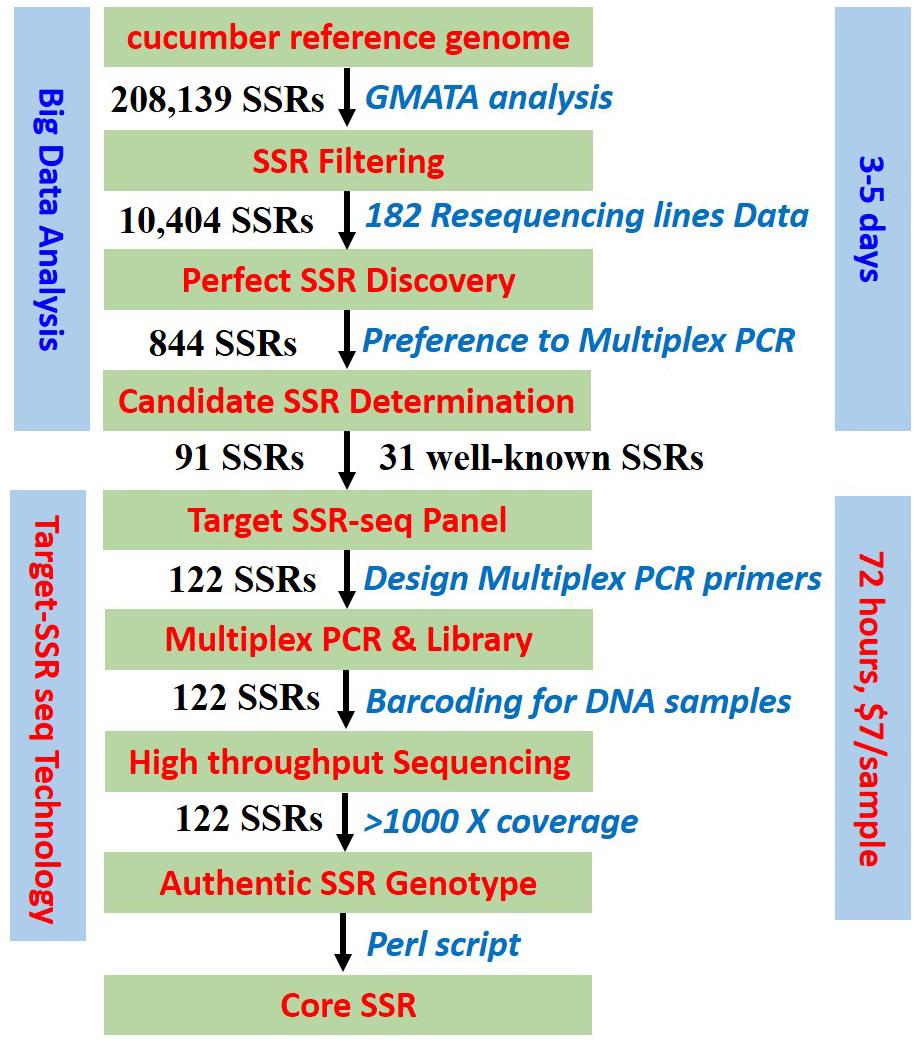
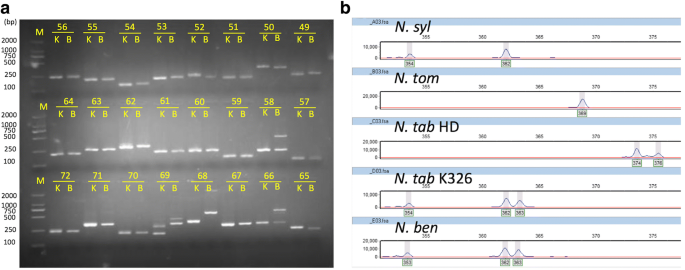
Frontiers | Target SSR-Seq: A Novel SSR Genotyping Technology Associate With Perfect SSRs in Genetic Analysis of Cucumber Varieties

Development of Genome-wide SSR Markers for Physical Map Construction with PCR-based Polymorphic SSRs in Jute (Corchorus Spp.) | SpringerLink

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect
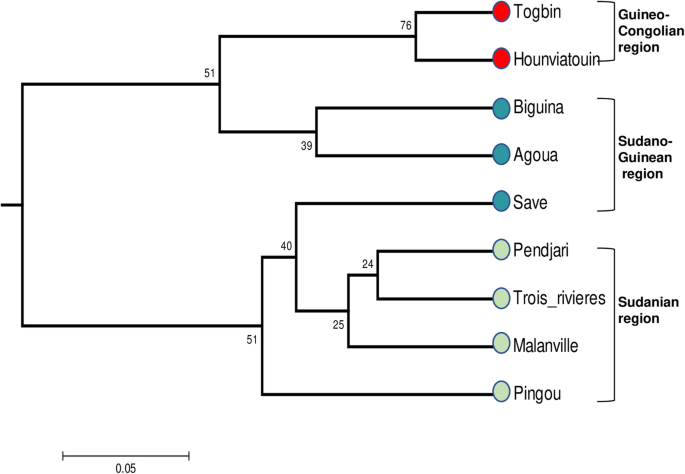
Transferability, development of simple sequence repeat (SSR) markers and application to the analysis of genetic diversity and population structure of the African fan palm (Borassus aethiopum Mart.) in Benin | BMC Genomic
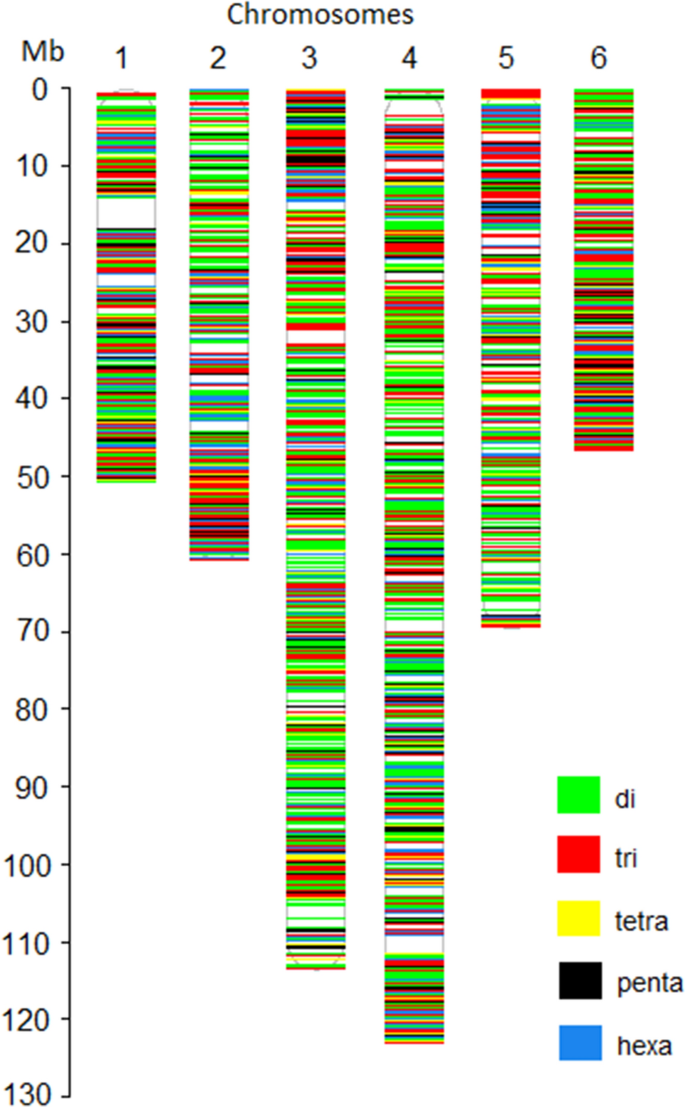
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Development and technical application of SSR-based individual identification system for Chamaecyparis taiwanensis against illegal logging convictions | Scientific Reports
High-Throughput Development of SSR Markers from Pea (Pisum sativum L.) Based on Next Generation Sequencing of a Purified Chinese Commercial Variety | PLOS ONE

PDF) A set of simple-sequence repeat (SSR) markers covering the Prunus genome | Elisabeth Dirlewanger - Academia.edu

Genes | Free Full-Text | Analysis of Genetic Diversity of Fescue Populations from the Highlands of Bolivia Using EST-SSR Markers

Management of a Walnut Germplasm Collection: Which of SSR or SNP Markers Are Most Suitable to Preserve Biodiversity? | bioRxiv

Data resolution (DR) curve for the 97 genomic SSR markers. The minimum... | Download Scientific Diagram
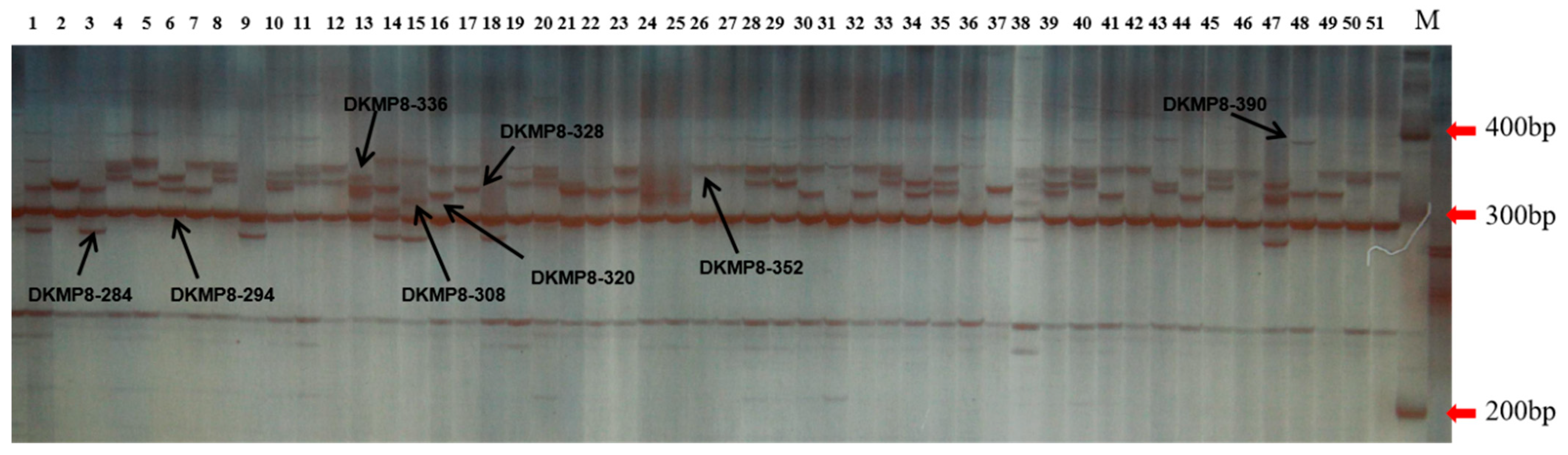
Forests | Free Full-Text | Genetic Diversity and Association Analysis among Germplasms of Diospyros kaki in Zhejiang Province Based on SSR Markers

Figure 2 - Identification of a minimal microsatellite marker panel for the fingerprinting of peach and nectarine cultivars
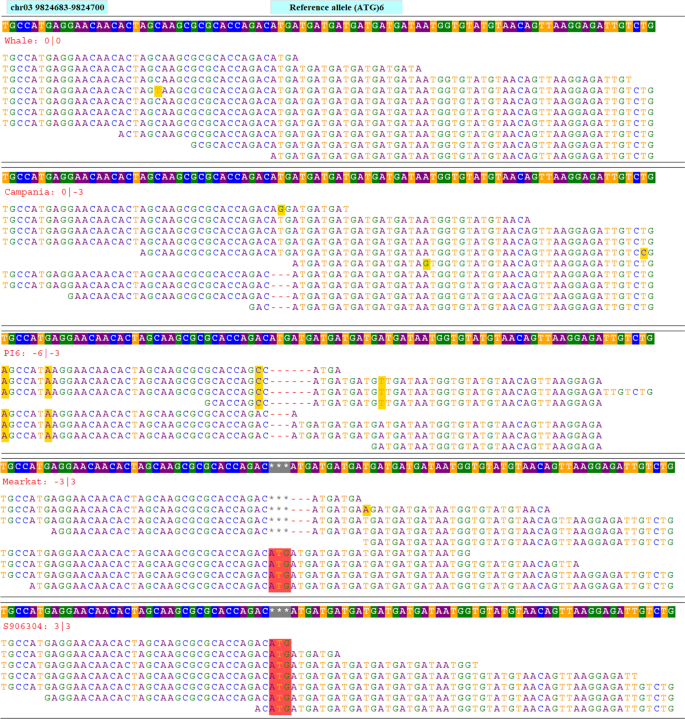
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
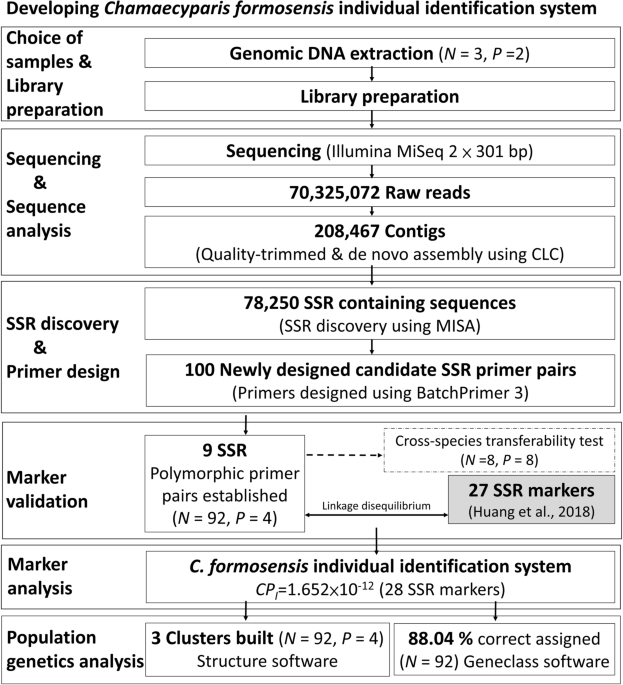
SSR individual identification system construction and population genetics analysis for Chamaecyparis formosensis | Scientific Reports
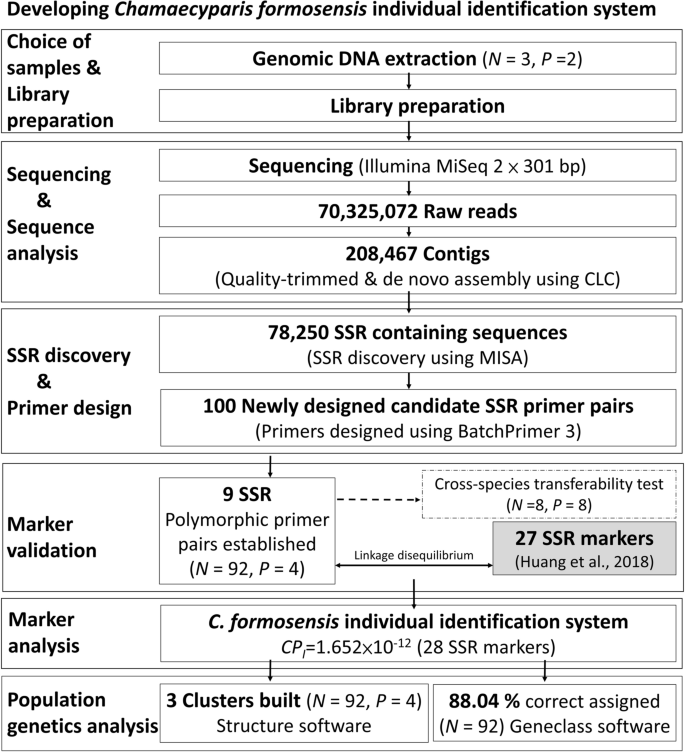
SSR individual identification system construction and population genetics analysis for Chamaecyparis formosensis | Scientific Reports

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
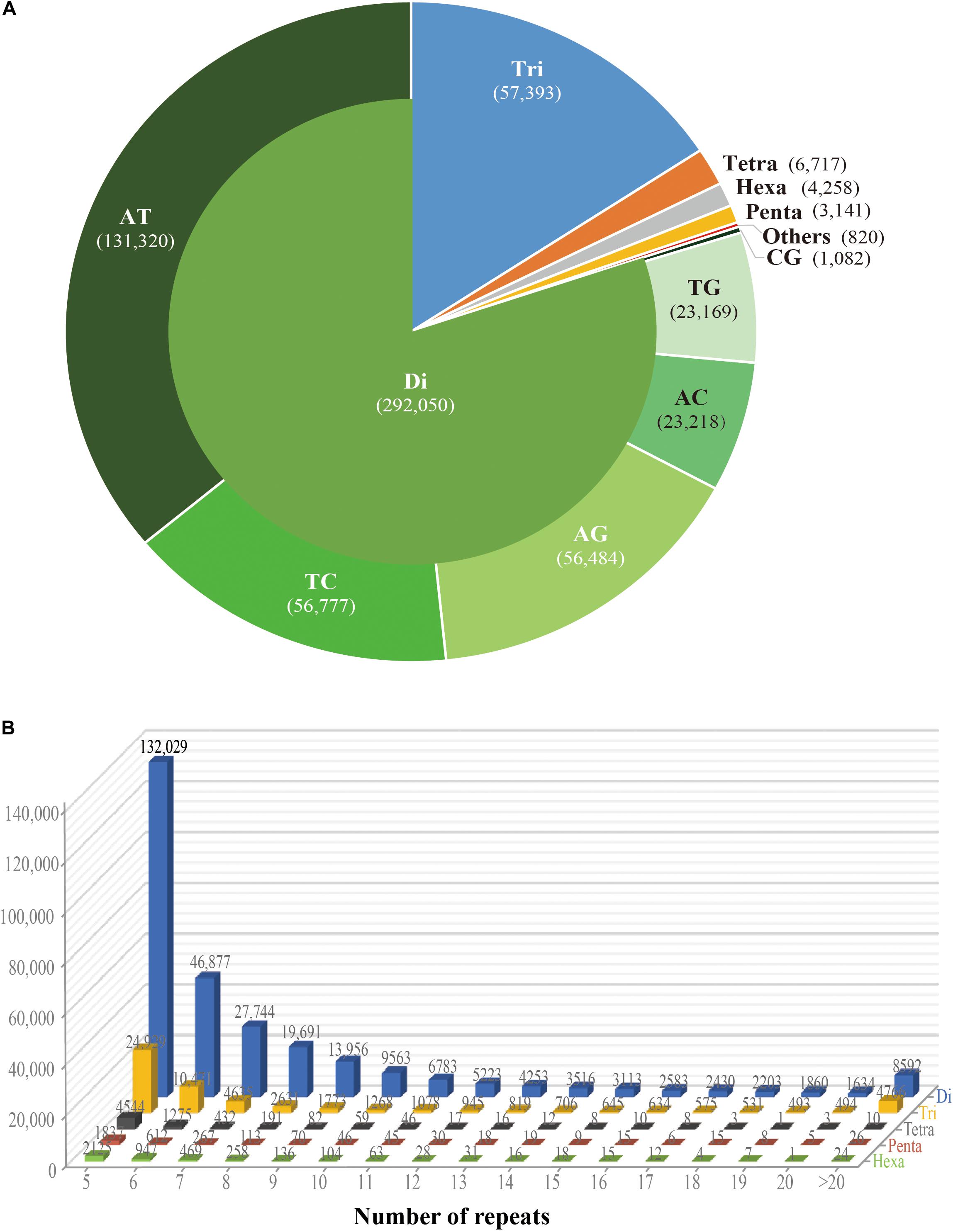
Frontiers | Development of a Large Gene-Associated SSR Marker Set and in-Depth Genetic Characterization in Scarlet Sage

Figure 1 | Association Analysis of SSR Markers with Phenology, Grain, and Stover-Yield Related Traits in Pearl Millet (Pennisetum glaucum (L.) R. Br.)